AZoOptics speaks to Prof. Simon Scheuring & Dr. Alma P. Perrino about their recent research using a new line-scanning high-speed atomic force microscopy technique. The method helps characterize the single-molecule kinetics of wild-type bR (bR-WT) exposed to continuous light and short light pulses.
How did you begin your research into the inner workings of light-sensing molecules?
The lab is specialized in atomic force microscopy (AFM)-based research of biological samples, with a particular interest in membrane phenomena. We develop and apply different high-speed AFM (HS-AFM) techniques to measure the dynamics of membrane proteins exposed to specific stimuli, such as force, environmental pH, ions or ligands, temperature, or light.
With the increasing promise of optogenetics, we thought that we could contribute an important piece of information – the single-molecule kinetics - to our understanding of the light-sensitive channels and pumps used in optogenetics. We became interested in studying proteins such as Channelrhodopsin-2 (ChR2), Halorhodopsin (NpHR) and Bacteriorhodopsin (bR), to better understand their photocycle. If we know more about their intrinsic function and structure, we are able to design new variants that are, for example, faster or more efficient at lower light intensity.
What is the importance of light-sensitive proteins and their study?
It was a major surprise when light-sensitive proteins were found in single-cell green algae. Up to then, these kinds of proteins were only found in the vision system of higher animals. However, with their discovery and the finding that some of them channeled or pumped ions, the promise was that if these proteins were inserted in animal neurons, they could work as cellular switches activated by light. Thus, light-sensitive proteins became the main tool in optogenetics.
What current challenges are there when studying the three-dimensional structure of bacteriorhodopsin?
The structure of bR is well known; the challenge that we addressed here was the study of its single-molecule kinetics. Most of the current spectroscopic methods provide information about the different states of the photocycle from ensemble-averaged data. At the single-molecule level, we can capture how fast an individual protomer in the purple membrane will respond to light stimulation, and when it could be activated again. HS-AFM had prior provided information about slow functioning mutants of bR, but studying the wild-type was elusive because its photocycle is too fast.
Can you explain your team’s research and the key findings?
In our work, we used high-speed atomic force microscopy methods to characterize the single-molecule kinetics of wild-type bR (bR-WT) exposed to continuous light and short light pulses.
We used a trick to boost the temporal resolution of HS-AFM: We first recorded images to target the proteins, and then changed the HS-AFM data acquisition to a mode that we term line-scanning. In this mode, instead of recording many scan lines to build an image, we record the same scan line many times and record the time evolution of the molecules in this scan line with almost 1000-fold increased temporal resolution. Thus, we recorded bR-WT conformational changes with millisecond temporal resolution and determined that the cytoplasmic gate opens 2.9 ms after photon absorption.
We also observed that a single protomer stays open on the cytoplasmic face for proton capture for 13.2 ms. Our experiments under continuous light and in light saturation regime revealed that a previously active protomer cannot be reactivated for another 37.6 ms. We propose that this lag time corresponds to the activity of the two neighboring protomers in the bR trefoil, ~32.2ms (2.9ms opening delay time + 13.2ms open state dwell time = 16.1ms x 2 neighboring protomers). This interpretation is in line with a crystallographic study that suggested that the enlarged open state could only occur in one protomer of three at any time. This means that the average maximum turnover that an individual protomer can reach is ~20 s−1.
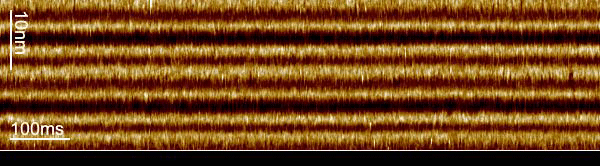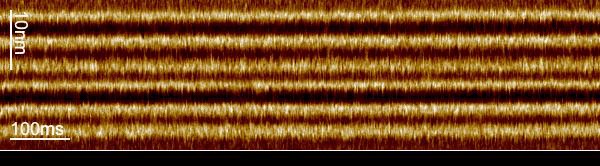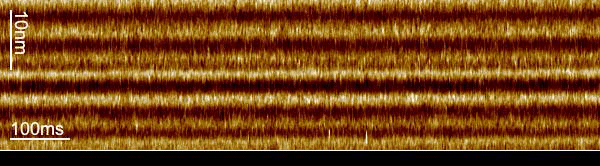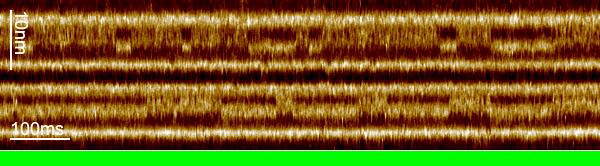
Image Credit: Weill Cornell Medicine
Your team has developed a new technique called line-scanning high-speed atomic force microscopy. Can you explain what this is and how it can be used to study light-sensitive proteins?
Recently, we introduced HS-AFM line scanning (HS-AFM-LS) for the characterization of GltPh, a member of the glutamate transporter family (Nat. Comm. 11, 5016 (2020)).
In this case, we studied how the transport domain moves vertically through the membrane. In HS-AFM-LS the y-scanner (also called slow scan-axis) is disabled, and the central x-scan line is repeatedly recorded.
The loss in the y-dimension comes with a strong increase, almost 1000-fold, in temporal resolution. For example, instead of taking 1 image with 600 scan lines per second, we record 600 times the same scan line per second, thus reaching a temporal resolution of 1.6 ms. From these measurements, profiles of the proteins located in this ‘slice’ of the image, we construct kymographs by stacking the scan lines as a function of time.
The increased temporal resolution allows us to resolve the bR-WT photocycle that is ~1000 times faster than that of slower mutants previously studied by HS-AFM regular imaging. From what we learned about the technique during our work on the glutamate transporter in 2020, we knew that we could apply it to bR, even though the displacement of the E-F loop is lateral instead of vertical and the speed of action much faster.
What are the positives of using this new atomic force microscopy technique?
First and most important, our HS-AFM line scanning method is a single molecule technique. Thus, we do not rely on ensemble averaging of data, but on single-molecule measurements that give us kinetic information about the behavior of individual protomers. It provides information on whether and when the same protomer can reactivate.
Second, the technique is label-free and carried out under physiological conditions. This gives us the freedom to choose pH, buffer, and type of light excitation, continuous light, or short light pulses. We are also watching the molecule itself and not a label that might have its own dynamics.
Finally, the resolution of the technique allows us to image with ~0.1 nm vertical and ~1 nm lateral imaging resolution, and – most importantly –, the technique provides easily here 1.6 ms temporal resolution, out beating most other single-molecule techniques.
What applications will this new technology be useful for?
We propose HS-AFM-LS as a method to study membrane protein single-molecule kinetics in general. The method reports Angstrom-range conformational dynamics at high temporal resolution, opening avenues to further application to other light-sensitive proteins or other dynamic membrane proteins, e.g. transporters, channels.
Is there a potential for this to be adopted on a global scale? What are the challenges that may be faced and how could these be overcome?
No, AFM is still a technique with low throughput. We are seeing constant advances that indicate things are moving in the right direction, in particular standardization of sample preparation and data analysis, as well as public code data repositories. But, in general, the AFM method is based on the physical contact between tip and sample and is not that prone for high throughput.
What are the next steps for the project?
Since we found such a surprising behavior of the trimer within the purple membrane, our short-term plan is to use other variants of bR, such as mutants bR-W12I or bR-F135I, that are unable to form lattices but show spectroscopically photocycle intermediates similar to the wild-type (~10 ms open state). This will help us to better understand the cooperativity and the mechanism of communication between protomers. A long-term plan is the application of this method to other light-sensitive proteins such as Channelrhodopsin-2 (ChR2) or Gloeobacter Rhodopsin (GR).
Where can readers find more information?
https://www.nature.com/articles/s41467-021-27580-2
https://laskerfoundation.org/winners/light-sensitive-microbial-proteins-optogenetics/
https://bio-afm-lab.com/
About Prof. Simon Scheuring
 Prof. Simon Scheuring is a Professor of Physiology and Biophysics in Anesthesiology at Weill Cornell Medicine, New York, USA. He is a trained biologist from the Biozentrum at the University of Basel, Switzerland.
Prof. Simon Scheuring is a Professor of Physiology and Biophysics in Anesthesiology at Weill Cornell Medicine, New York, USA. He is a trained biologist from the Biozentrum at the University of Basel, Switzerland.
Simon Scheuring has been awarded an INSERM Avenir (2005), the médaille de la Ville de Paris (2007), a European Research Council (ERC) consolidator grant (2012), the grand prix Robert Debré (2013), and the NIH director’s pioneer award (2019).
Scheuring's objective is to head a dynamic research team with members from different scientific fields ranging from biology to physics and engineering to develop and use atomic force microscopy-based technologies to provide novel insights into the processes taking place in biological membranes.
About Dr. Alma P. Perrino
 Dr. Alma P. Perrino is a postdoctoral researcher at the Department of Anesthesiology at Weill Cornell Medicine, New York, USA, in the laboratory of Prof. Simon Scheuring.
Dr. Alma P. Perrino is a postdoctoral researcher at the Department of Anesthesiology at Weill Cornell Medicine, New York, USA, in the laboratory of Prof. Simon Scheuring.
Alma got her PhD in Physics from Universidad Autónoma de Madrid, Spain. She develops and applies new high-speed atomic force microscopy (HS-AFM) methods for the study of the dynamics and structure of membrane proteins.
Disclaimer: The views expressed here are those of the interviewee and do not necessarily represent the views of AZoM.com Limited (T/A) AZoNetwork, the owner and operator of this website. This disclaimer forms part of the Terms and Conditions of use of this website.