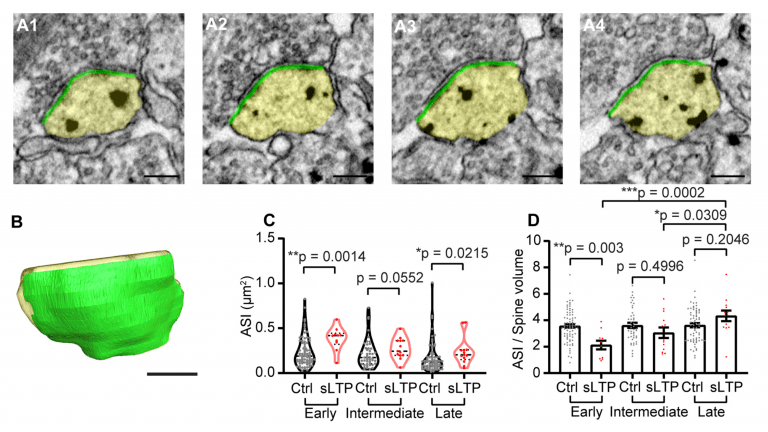
Image Credit: Max Planck Florida Institute for Neuroscience
Neuroscientists perform a very similar task. They have vigorously attempted to improve methods to capture perfect, crystal-clear images. Instead of picturesque outdoor sites, neuroscientists take detailed snapshots of brain cells and their small-scale structures.
In this process, the brain cells can actively strengthen key connections and weaken those that are less required. This process is considered to underlie the way it is understood and remembered. However, uncovering the fine structure of the spine in detail during a dynamic process is a challenging task. Until now, imaging methodologies cannot do so.
Reportedly, the researchers at the Yasuda Lab have designed a new robust imaging technique, with the ability to visualize the fine, ultrastructural alteration to dendritic spines during structural plasticity. The study was published in The Journal of Neuroscience.
The researchers tweaked correlative light and electron microscopy (CLEM) and developed a new imaging method to obtain the best that both imaging modalities can offer.
Dendritic spines are such small-scale neuronal compartments, that it’s difficult to get an accurate picture of what’s actually occurring in terms of structural changes using traditional imaging methods. Using more standard optical techniques like 2-photon microscopy, dendritic spines look like smooth spheres.
Dr. Ryohei Yasuda, Scientific Director, Max Planck Florida Institute for Neuroscience
“So, we were interested in learning what changes occur during the various stages of structural plasticity, at a resolution where we could take a deeper look at the spine’s complexity. In actuality, we know from using more powerful imaging methods, like electron microscopy, that the actual size and shape of spines are far more complex,” Dr. Yasuda added.
First, the researchers induced structural plasticity in a single dendritic spine by employing 2-photon optical microscopy and glutamate uncaging. This induced spine was later fixed in time at one of the three unique timepoints. This represented the important level of structural plasticity.
In association with the MPFI Electron Microscopy (EM) Core, brain tissue samples with the stimulated spines were cut into ultrathin sections by employing a specialized device known as ATUMtome. Then, the sections were re-imaged using the extreme resolving power of the Electron Microscope to uncover the ultrastructural details and rebuild precise pictures of the complex topology of the spine.
When we started this project, our goal was to see if it was even possible to collect spines at various stages of structural plasticity, successfully relocate them, and resolve their ultrastructure using EM. Single, spine-specific forms of structural plasticity have never been imaged in this way before. Dr. Naomi kamasawa, Head of MPFI’s EM Core, was instrumental in helping to establish and optimize our EM workflow for the project.
Ye Sun, PhD, Study First Author and Former Graduate Student, Yasuda Lab, Max Planck Florida Institute for Neuroscience
While analyzing the reconstructed spine images, the researchers observed distinct changes to a protein-rich region of dendritic spines, known as postsynaptic density (PSD). This region is crucial for the spine and known for regulating synaptic strength and plasticity.
The MPFI researchers revealed that when compared to control spines, the area and size of the PSD region were notably greater in spines that underwent structural plasticity. The growth of PSD in these spines has happened over a slower timescale, requiring hours to attain its maximum change.
Fascinatingly, even when the growth was at a slow rate, the PSD structure in the simulated spines reorganized faster. Following the induction of structural plasticity, there was a quick increase in PSD complexity, causing a remarkable transformation in the shape and structural features.
Our imaging strategy synergizes the best of both optical and EM microscopies, allowing us to study spine structural changes never before seen in nanoscale resolution. For the future, our lab is interested in using this new protocol in combination with advanced molecular techniques, such as SLENDR, to study individual protein dynamics in tandem with finely detailed structural changes during spine structural plasticity.
Dr. Ryohei Yasuda, Scientific Director, Max Planck Florida Institute for Neuroscience
The research was supported by the National Institutes of Health Grants, the Brain Research Foundation and the Max Planck Florida Institute for Neuroscience.
Journal Reference:
Sun, Y., et al. (2021) Rapid Ultrastructural Changes in the PSD and Surrounding Membrane after Induction of Structural LTP in Single Dendritic Spines. Journal of Neuroscience. doi.org/10.1523/JNEUROSCI.1964-20.2021.